arrghh
- Posted by darkeye on March 19th, 2008 filed in experiments, log
- 1 Comment »
Today was the day to see if indeed we can turn on and off the level of fluorescence in our bacteria by changing the acidity of the surrounding environment. Well, in short, the results are not really good. But, there’s always hope, in this case brought by Jos Arents from the University of Amsterdam 🙂
At first, we had a hard time trying to find a light with the right excitation wavelength for our cherished green fluorescent protein. After experimenting with various UV lights of 245 nm and 365 nm wavelengths, we also tried out a disco-proof UV light (no, really, that’s also 365 nm actually). The pGLO strain is as fluorescent as it can get, but the pH-sensitive GFP is not really shining. Looking into the article mentioning the TorA-GFPmut3* GFP, it seems that the right excitation wavelength is about 500 nm – which is actually in the visible light spectrum, and is green. But – having found a lamp of 505 nm, there’s no visible fluorescence coming out of it. Even though that would be around 545 nm, quite close, between green and yellow, one should be able to see a visible level of different light than the excitation light.
So it seems there’s a general problem with the fluorescent protein in our bacteria strain. Tomorrow I’ll have a chance to run it through a machine that will give me an excitation curve, and see really if it is fluorescent and how.
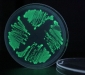
Anyway, even with limited level of visible fluorescence, we tried out dropping some acid (HCl) on the agar plate with the bacteria. Well, you can see a minimal decrease in fluorescence, but I’m not sure if this is just a reaction of the agar base, or there’s really something happening to the fluorescence of the bacteria. Moreover, the fluorescence level does not seem to return to the previous state, which it should in about 20 minutes.
Just out of curiosity, I also put some acid on the pGLO bacteria, which indeed makes it un-fluorescent, but I think it kills it right away as well. The bacteria turns white-ish under normal light, and the level of fluorescence does not return to normal here either.
Meanwhile, Jos Arents at the University of Amsterdam has succeeded in his transformation, and made two plates to purify the results. He grew the bacteria in LB with ampicillin, chloroamphenicol and kanamycin, so that only the desired bacteria survive. And thus he put them into three different environments:
- nothing (just the standard plate)
- IPTG
- arabinose and IPTG
The expectation is that the above will produce, correspondingily:
- nothing really (plain bacteria)
- pHluorin but no proteorhodopsin (pH-sensitive GFP, but not light-sensitive parts)
- both pHluorin and proteorhodopsin (both pH-sensitive GFP and light-sensitive parts that change the pH)
Tomorrow, he’ll see the results. I hope to be there…



March 19th, 2008 at 12:44 pm
“we tried out dropping some acid”…. oh, the wonders of a bio experiments in amsterdam 🙂