the future doesn’t glow as bright
- Posted by darkeye on March 14th, 2008 filed in experiments, log
- 2 Comments »
Today there was a chance to take a look at the new growth of both E. Coli strains, one that has the gGLO DNA in it from BIO-RAD, the other having the TorA-GFPmut3* GFP transformed into it. The latter is the one that should have a level of fluorescence correlating to the pH value around it.
The sad news is, that the pGLO bacteria, which was extremely bright after performing the original transformation, doesn’t really glow now at all. The bacteria with the TorA-GFPmut3* GFP in itdoes glow, but not really bright. And it seems it has a tendency to glow when lit with lower wavelength UV light – which is dangerous to the bacteria itself an to humans as well.
Anyway, we put the TorA-GFPmut3* GFP version into deep freeze storage (-70 °C), under the code MS 903, so that it is readily available whenever needed.
So what could have gotten wrong with the pGLO bacteria? Don’t really know – maybe there’s too little arabinose in the agar gel we’ve put the bacteria on. Arabinose is the thing triggering the specific DNA to express the fluorescent protein, so if it’s not enough, we won’t see a lot of glow.
As for the TorA-GFPmut3* GFP version, the fact that it has a characteristic of liking lower wavelength UV light doesn’t seem to be nice. Also, the brightness of the glow is not really visible in daylight conditions, which is not good from an installation perspective.
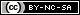
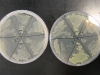
For the experiment, we made a petri dish with six sections, containing three different kinds of bacteria. Each are placed twice, opposite to each other, marked 1 and 2. The markings are:
- GFP: the bacteria containing the TorA-GFPmut3* GFP
- GLO: the bacteria containing pGLO
- -: the bacteria not containing fluorescent protein.
The dish on the left does not have arabinose in it, thus the fluorescent proteins should not be expressed in it. The plate on the right has arabinose, thus the GFP and GLO sections should be fluorescent:
The strange thing is, the bacteria with TorA-GFPmut3* GFP doesn’t glow at all with the UV LED that was supplied with the BIO-RAD kit. Thus we had to look for different UV lamps. Good thing about a considerably sized university is that there is a lot of cool hardware around.
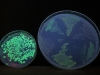
Here’s a picture of the level of fluorescence when the excitation wavelength is 254 nm. The small dish on the left is a growth from the original pGLO experiment. The big dish contains six sections, which are three different bacteria strains (see above):
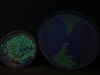
And here’s a picture when the excitation is done with a light of 365 nm, having taken the picture with the same camera settings. These guys are visibly less excited by this light:
So what we did, is try to grow the pGLO bacteria again, now taking samples from the plate that included the arabinose in the first place. After all, these are the ones that are very visibly fluorescent. We’ll see by Monday if this works out.
On the other hand, we could try to grow both of them with a lot more arabinose around, and see what happens.
Next week, we’ll try to verify if there’s a visible level of change in fluorescence in the bacteria containing TorA-GFPmut3* GFP. For that, we’ll grow even lawns of bacteria, and then try to influence the pH value around. Another suggestion was that we should grow them on a liquid base, where one can measure the pH value very precisely, and see if there’s a change in fluorescence there.
It will be also worthwhile to look around for other GFPs that change their level of fluorescence when the surroinding pH changes – maybe there are some with different excitation characteristics – that is, they are excited more by near-human-visible light, and glow also brighter.
2 Responses to “the future doesn’t glow as bright”
Leave a Comment
You must be logged in to post a comment.








March 14th, 2008 at 3:44 pm
What about the pH of our plates? Could we be in the low-fluorescence range? Or should the intracellular pH really be totally independent of that?
March 15th, 2008 at 3:10 am
good point – but according to the Wilks article, the intracellular pH is maintained at around 7 by E. Coli on the long run. when the external pH changes to about 5.5, the internal pH first drops (in 4 seconds), then is raised back to about 7 in the order of 10 – 20 minutes.